Code
| age | chol | target | |
|---|---|---|---|
| 527 | 62 | 209 | 1 |
| 359 | 53 | 216 | 1 |
| 447 | 55 | 289 | 0 |
CSCI-866-001: Data Mining & Knowledge Discovery



OneHotEncoding.import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.preprocessing import OneHotEncoder
data = pd.read_csv(path + "/heart.csv")
quan_vars = ['age','trestbps','chol','thalach','oldpeak']
qual_vars = ['sex','cp','fbs','restecg','exang','slope','ca','thal','target']
# Convert to correct types
for i in quan_vars:
data[i] = data[i].astype('float')
for i in qual_vars:
data[i] = data[i].astype('category')
data.drop_duplicates(inplace=True)
# Train test split
from sklearn.model_selection import train_test_split
X, y = data.iloc[:,:-1], data.iloc[:,-1]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, stratify=y, random_state=42)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
encoder = OneHotEncoder(drop='first', sparse_output=False)
# OnehotEncoding and Scaling
X_train_cat = encoder.fit_transform(X_train.select_dtypes(include="category"))
X_train_encoded = scaler.fit_transform(np.column_stack([X_train.select_dtypes(include="number").to_numpy(), X_train_cat]))
X_test_cat = encoder.transform(X_test.select_dtypes(include="category"))
X_test_encoded = scaler.transform(np.column_stack([X_test.select_dtypes(include="number").to_numpy(), X_test_cat]))
# KNN
from sklearn.svm import SVC
svm_model = SVC(kernel='linear', random_state=42)
# Define the hyperparameter grid
param_grid = {'C': [0.1, 0.2, 0.25, 0.3, 0.4, 0.5, 0.75, 1]}
# Perform GridSearch for optimal C
grid_search = GridSearchCV(svm_model, param_grid, cv=20, scoring='accuracy')
grid_search.fit(X_train_encoded, y_train)
# Get the best parameter
best_C = grid_search.best_params_['C']
# Train final model using best C
final_model = SVC(kernel='rbf', C=best_C, random_state=42)
final_model.fit(X_train_encoded, y_train)
# Predict on test set
y_pred = final_model.predict(X_test_encoded)
from sklearn.metrics import roc_auc_score, accuracy_score, precision_score, recall_score, f1_score, confusion_matrix, ConfusionMatrixDisplay
test_perf = pd.DataFrame(
data={'Accuracy': accuracy_score(y_test, y_pred),
'Precision': precision_score(y_test, y_pred),
'Recall': recall_score(y_test, y_pred),
'F1-score': f1_score(y_test, y_pred),
'AUC': roc_auc_score(y_test, y_pred)},
columns=["Accuracy", "Precision", "Recall", "F1-score", "AUC"],
index=["LSVM"])
test_perf| Accuracy | Precision | Recall | F1-score | AUC | |
|---|---|---|---|---|---|
| LSVM | 0.852459 | 0.852941 | 0.878788 | 0.865672 | 0.850108 |
import numpy as np
import numpy as np
import plotly.graph_objects as go
import plotly.express as px
from plotly.subplots import make_subplots
from sklearn.datasets import make_classification, make_circles, make_moons
# Generate 2D binary classification datasets
def generate_2d_dataset(n_samples=500, boundary_type='linear', noise=0.1, random_state=42, mis_class = 0.1):
"""
Generate a 2D binary classification dataset with linear or non-linear decision boundary.
Parameters:
-----------
n_samples : int, default=500
Number of samples to generate
boundary_type : str, default='linear'
Type of decision boundary: 'linear', 'circular', 'moons', 'spiral'
noise : float, default=0.05
Amount of noise to add to the data (0.0 to 1.0)
random_state : int, default=42
Random state for reproducibility
Returns:
--------
X : ndarray of shape (n_samples, 2)
Feature matrix
y : ndarray of shape (n_samples,)
Target labels (0 or 1)
"""
np.random.seed(random_state)
if boundary_type == 'linear':
# Generate well-separated linearly separable data
X, y = make_classification(
n_samples=n_samples,
n_features=2,
n_redundant=0,
n_informative=2,
n_clusters_per_class=1,
random_state=random_state,
class_sep=1.5, # Increased separation
flip_y = mis_class
)
# Add minimal noise
X += np.random.normal(0, noise, X.shape)
elif boundary_type == 'circular':
# Generate well-separated circular decision boundary
X, y = make_circles(
n_samples=n_samples,
noise=noise, # Reduced noise
factor=0.4, # Better separation between circles
random_state=random_state
)
elif boundary_type == 'moons':
# Generate well-separated moon-shaped decision boundary
X, y = make_moons(
n_samples=n_samples,
noise=noise, # Reduced noise
random_state=random_state
)
elif boundary_type == 'spiral':
# Generate better separated spiral decision boundary
n_per_class = n_samples // 2
theta = np.linspace(0, 3*np.pi, n_per_class) # Reduced spiral length
# First spiral (class 0)
r1 = theta / (1.5*np.pi) # Slower growth rate
x1 = r1 * np.cos(theta)
y1 = r1 * np.sin(theta)
# Second spiral (class 1) - better separated
x2 = r1 * np.cos(theta + np.pi)
y2 = r1 * np.sin(theta + np.pi)
# Combine data
X = np.vstack([np.column_stack([x1, y1]), np.column_stack([x2, y2])])
y = np.hstack([np.zeros(n_per_class), np.ones(n_per_class)])
# Add minimal noise
X += np.random.normal(0, noise * 0.3, X.shape)
# Shuffle the data
indices = np.random.permutation(len(X))
X, y = X[indices], y[indices]
else:
raise ValueError("boundary_type must be one of: 'linear', 'circular', 'moons', 'spiral'")
return X, y.astype(int)
#| echo: true
#| code-fold: true
import warnings
warnings.filterwarnings('ignore')
# Plot decision boundary and data
def plot_decision_boundary(X, y, model=None, title="Decision Boundary",
resolution=100, alpha_contour=0.7, alpha_points=0.8,
colorscale='RdYlBu', point_size=8, show_mesh=True):
"""
Create an interactive decision boundary plot using Plotly with color gradients.
Parameters:
-----------
X : array-like of shape (n_samples, 2)
Feature matrix (must be 2D)
y : array-like of shape (n_samples,)
Target labels
model : sklearn estimator or None, default=None
Trained model that implements predict() and predict_proba() or decision_function().
If None, only plots the dataset without decision boundary.
title : str, default="Decision Boundary"
Title for the plot
resolution : int, default=100
Resolution of the decision boundary mesh
alpha_contour : float, default=0.7
Transparency of the decision boundary
alpha_points : float, default=0.8
Transparency of the data points
colorscale : str, default='RdYlBu'
Plotly colorscale for the decision boundary
point_size : int, default=8
Size of the scatter points
show_mesh : bool, default=True
Whether to show the decision boundary mesh (ignored if model is None)
Returns:
--------
fig : plotly.graph_objects.Figure
Interactive Plotly figure
"""
# Create the figure
fig = go.Figure()
# Add decision boundary contour only if model is provided and show_mesh is True
if model is not None and show_mesh:
# Ensure model is fitted
if not hasattr(model, 'predict'):
raise ValueError("Model must be fitted and have a predict method")
# Create mesh grid
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, resolution),
np.linspace(y_min, y_max, resolution))
mesh_points = np.c_[xx.ravel(), yy.ravel()]
# Get predictions for the mesh
try:
# Try to get probability predictions for smoother boundaries
if hasattr(model, 'predict_proba'):
Z = model.predict_proba(mesh_points)[:, 1] # Probability of class 1
elif hasattr(model, 'decision_function'):
Z = model.decision_function(mesh_points)
# Normalize decision function output to [0, 1] range
Z = (Z - Z.min()) / (Z.max() - Z.min())
else:
Z = model.predict(mesh_points).astype(float)
except Exception as e:
print(f"Error getting model predictions: {e}")
Z = model.predict(mesh_points).astype(float)
Z = Z.reshape(xx.shape)
# Add decision boundary contour
fig.add_trace(go.Contour(
x=np.linspace(x_min, x_max, resolution),
y=np.linspace(y_min, y_max, resolution),
z=Z,
colorscale=colorscale,
opacity=alpha_contour,
showscale=True,
colorbar=dict(
title="Decision<br>Confidence",
tickmode="linear",
tick0=0,
dtick=0.2
),
contours=dict(
start=0,
end=1,
size=0.1,
),
name="Decision Boundary"
))
# Add data points
unique_labels = np.unique(y)
colors = px.colors.qualitative.Set1[:len(unique_labels)]
for i, label in enumerate(unique_labels):
mask = y == label
fig.add_trace(go.Scatter(
x=X[mask, 0],
y=X[mask, 1],
mode='markers',
marker=dict(
size=point_size,
color=colors[i],
opacity=alpha_points,
line=dict(width=1, color='black')
),
name=f'Class {label}',
hovertemplate=f'<b>Class {label}</b><br>' +
'Feature 1: %{x:.2f}<br>' +
'Feature 2: %{y:.2f}<br>' +
'<extra></extra>'
))
# Update layout
fig.update_layout(
title=dict(
text=title,
x=0.5,
font=dict(size=18, family="Arial Black")
),
xaxis_title="Feature 1",
yaxis_title="Feature 2",
width=700,
height=600,
showlegend=True,
legend=dict(
yanchor="top",
y=0.99,
xanchor="left",
x=0.01,
bgcolor="rgba(255,255,255,0.8)",
bordercolor="black",
borderwidth=1
),
plot_bgcolor='white',
paper_bgcolor='white'
)
# Make axes equal
fig.update_xaxes(scaleanchor="y", scaleratio=1)
fig.update_yaxes(scaleanchor="x", scaleratio=1)
# Add information text if no model is provided
if model is None:
fig.add_annotation(
xref="paper", yref="paper",
x=0.02, y=0.02,
text="Dataset visualization<br>(No model provided)",
showarrow=False,
font=dict(size=12, color="gray"),
bgcolor="rgba(255,255,255,0.8)",
bordercolor="gray",
borderwidth=1
)
return fig
n = 300
data1 = generate_2d_dataset(n_samples=n, boundary_type="moons", noise=0.2)
fig1 = plot_decision_boundary(data1[0], data1[1], model=None)
fig1.update_layout(
title="Non-linear Decision Boundary Example (Moons Dataset)",
width=500,
height=270
)
fig1.show()# Input / output data
X1, y1 = data1[0], data1[1]
# train and test split
from sklearn.model_selection import train_test_split
X_train1, X_test1, y_train1, y_test1 = train_test_split(X1, y1, test_size=0.2, random_state=42)
# Build SVM model
from sklearn.svm import SVC
svc1 = SVC(kernel='linear') # initialization of the model
svc1 = svc1.fit(X_train1, y_train1) # Train the model
fig_bdr1 = plot_decision_boundary(X_test1, y_test1, model=svc1)
fig_bdr1.update_layout(
title="Linear Decision Boundary by SVM",
width=500,
height=270
)
fig_bdr1.show()svc2 = SVC(kernel='rbf', random_state=42) # initialization of the model
svc2 = svc2.fit(X_train1, y_train1) # Train the model
fig_bdr2 = plot_decision_boundary(X_test1, y_test1, model=svc2)
fig_bdr2.update_layout(
title="Non-linear Decision Boundary by SVM (RBF Kernel)",
width=500,
height=270
)
fig_bdr2.show()svc3 = SVC(kernel='poly', degree=3, random_state=42) # initialization of the model
svc3 = svc3.fit(X_train1, y_train1) # Train the model
fig_bdr3 = plot_decision_boundary(X_test1, y_test1, model=svc3)
fig_bdr3.update_layout(
title="Non-linear Decision Boundary by SVM (Polynomial-3 Kernel)",
width=500,
height=270
)
fig_bdr3.show()svm_model = SVC(kernel='rbf', random_state=42)
# Define the hyperparameter grid
param_grid = {'C': [5, 6, 7, 8, 10, 12.5, 15, 17, 20, 30, 45, 50]}
# Perform GridSearch for optimal C
grid_search = GridSearchCV(svm_model, param_grid, cv=20, scoring='accuracy')
grid_search.fit(X_train_encoded, y_train)
# Get the best parameter
best_C_rbf = grid_search.best_params_['C']
# Train final model using best C
final_model = SVC(kernel='rbf', C=best_C_rbf, random_state=42)
final_model.fit(X_train_encoded, y_train)
# Predict on test set
y_pred = final_model.predict(X_test_encoded)
from sklearn.metrics import roc_auc_score, accuracy_score, precision_score, recall_score, f1_score, confusion_matrix, ConfusionMatrixDisplay
test_perf = pd.concat([test_perf, pd.DataFrame(
data={'Accuracy': accuracy_score(y_test, y_pred),
'Precision': precision_score(y_test, y_pred),
'Recall': recall_score(y_test, y_pred),
'F1-score': f1_score(y_test, y_pred),
'AUC': roc_auc_score(y_test, y_pred)},
columns=["Accuracy", "Precision", "Recall", "F1-score", "AUC"],
index=["RBF-SVM"])])
# Polynomial 3 kernel
svm_model = SVC(kernel='poly', degree=3, random_state=42)
# Perform GridSearch for optimal C
grid_search = GridSearchCV(svm_model, param_grid, cv=5, scoring='accuracy')
grid_search.fit(X_train_encoded, y_train)
# Get the best parameter
best_C_poly = grid_search.best_params_['C']
# Train final model using best C
final_model = SVC(kernel='poly', degree=3, C=best_C_poly, random_state=42)
final_model.fit(X_train_encoded, y_train)
# Predict on test set
y_pred = final_model.predict(X_test_encoded)
from sklearn.metrics import roc_auc_score, accuracy_score, precision_score, recall_score, f1_score, confusion_matrix, ConfusionMatrixDisplay
test_perf = pd.concat([test_perf, pd.DataFrame(
data={'Accuracy': accuracy_score(y_test, y_pred),
'Precision': precision_score(y_test, y_pred),
'Recall': recall_score(y_test, y_pred),
'F1-score': f1_score(y_test, y_pred),
'AUC': roc_auc_score(y_test, y_pred)},
columns=["Accuracy", "Precision", "Recall", "F1-score", "AUC"],
index=["Poly3-SVM"])])
test_perf| Accuracy | Precision | Recall | F1-score | AUC | |
|---|---|---|---|---|---|
| LSVM | 0.852459 | 0.852941 | 0.878788 | 0.865672 | 0.850108 |
| RBF-SVM | 0.819672 | 0.866667 | 0.787879 | 0.825397 | 0.822511 |
| Poly3-SVM | 0.852459 | 0.875000 | 0.848485 | 0.861538 | 0.852814 |
rbf’, ‘poly’…OneHotEncoderStandardScaler, MinMaxScaler…for t = 1,2,..,T:
Prediction:
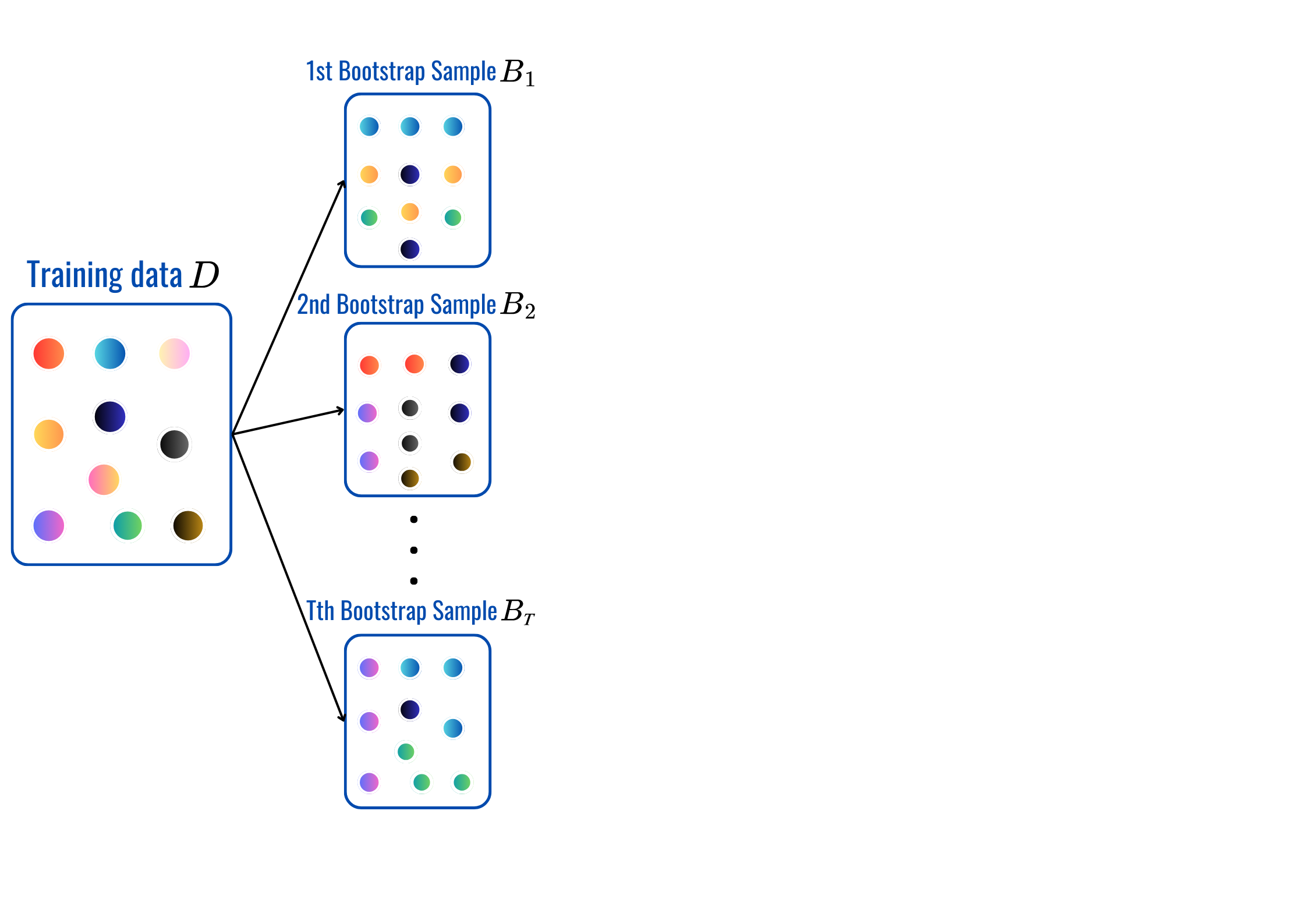
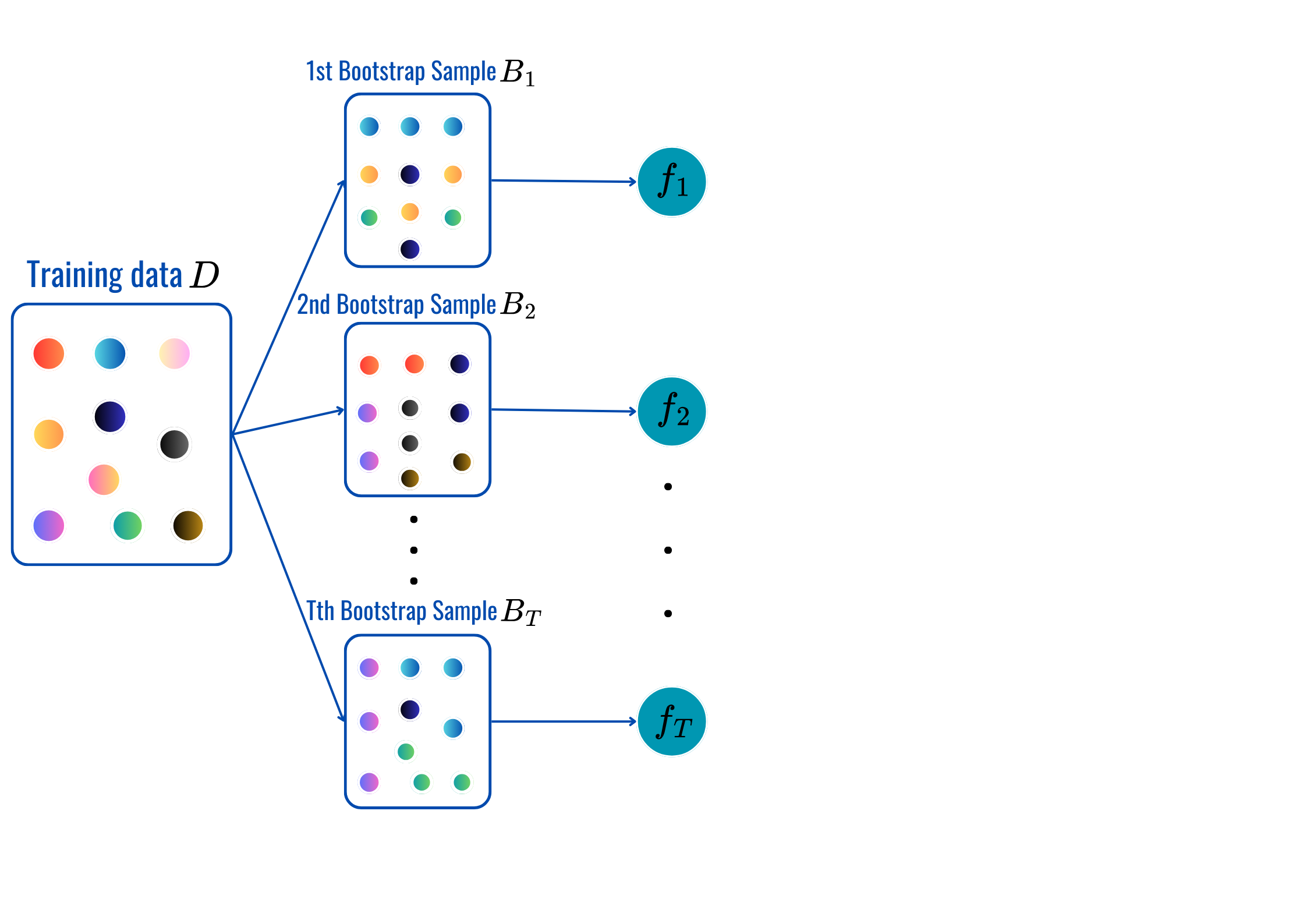
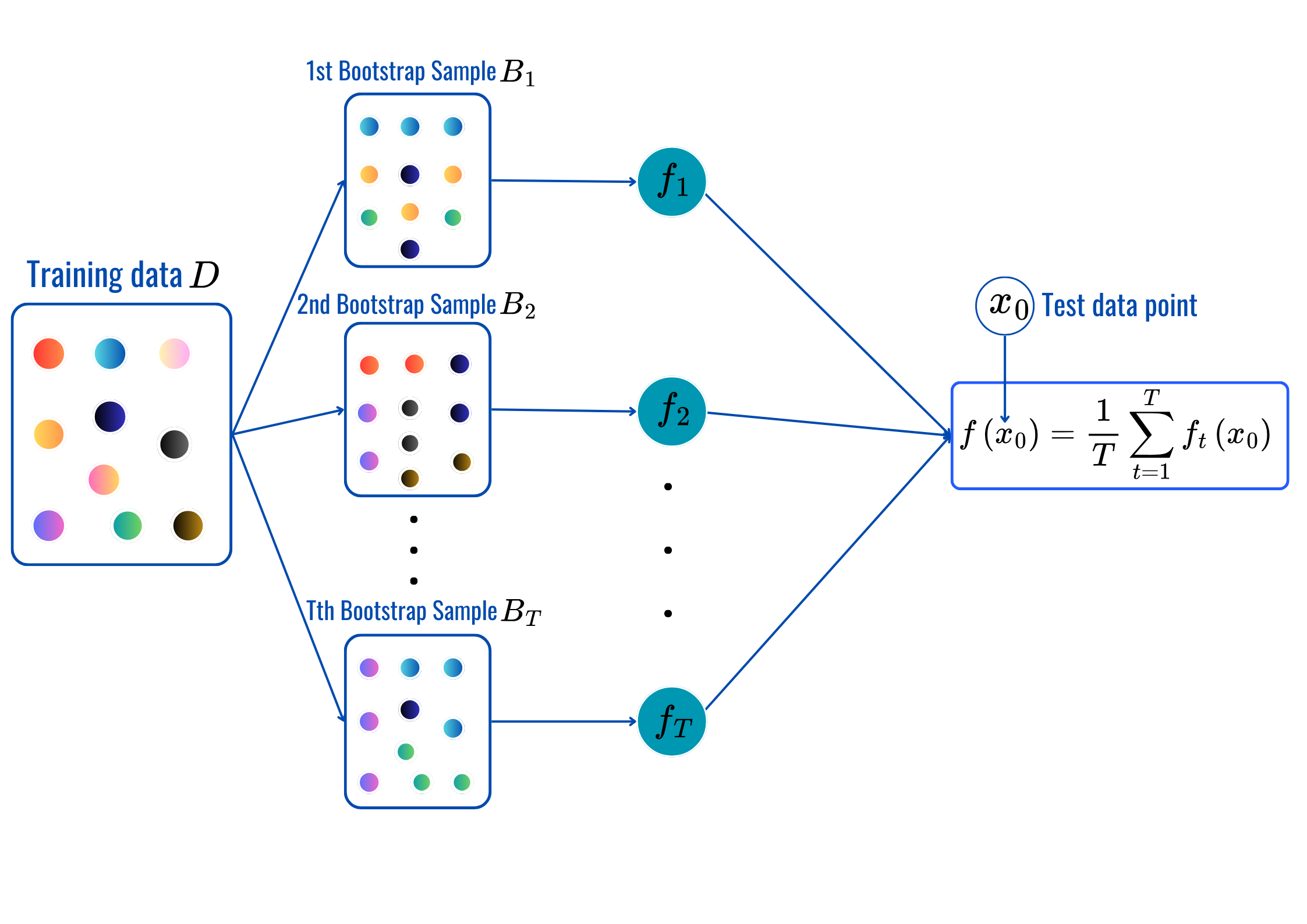
max_depth, min_sample_split, min_samples_leaf and citerion.from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
# Define base Decision Tree (with tunable parameters)
base_tree = DecisionTreeClassifier()
# Set up Bagging Classifier
bagging_model = BaggingClassifier(estimator=base_tree)
# Define hyperparameter grid
param_grid = {
'n_estimators': [50, 100, 200, 300, 500], # Number of trees in Bagging
'max_samples': [0.1, 0.25, 0.4, 0.5, 0.6], # Proportion of samples per tree
'estimator__min_samples_leaf': [3, 5, 7, 10], # Minimum samples per leaf
'estimator__criterion': ["gini", "entropy"], # Splitting criterion
}
grid_search = GridSearchCV(bagging_model, param_grid, cv=5, n_jobs=-1, scoring='accuracy')
grid_search.fit(X_train_encoded, y_train)
# Best parameters and corresponding model
best_model = grid_search.best_estimator_
print(f"Best parameters: {grid_search.best_params_}")
# Make predictions with the best model
y_pred = best_model.predict(X_test_encoded)
test_perf = pd.concat([test_perf, pd.DataFrame(
data={'Accuracy': accuracy_score(y_test, y_pred),
'Precision': precision_score(y_test, y_pred),
'Recall': recall_score(y_test, y_pred),
'F1-score': f1_score(y_test, y_pred),
'AUC': roc_auc_score(y_test, y_pred)},
columns=["Accuracy", "Precision", "Recall", "F1-score", "AUC"],
index=["Bagging"])])
test_perfBest parameters: {'estimator__criterion': 'entropy', 'estimator__min_samples_leaf': 3, 'max_samples': 0.4, 'n_estimators': 50}| Accuracy | Precision | Recall | F1-score | AUC | |
|---|---|---|---|---|---|
| LSVM | 0.852459 | 0.852941 | 0.878788 | 0.865672 | 0.850108 |
| RBF-SVM | 0.819672 | 0.866667 | 0.787879 | 0.825397 | 0.822511 |
| Poly3-SVM | 0.852459 | 0.875000 | 0.848485 | 0.861538 | 0.852814 |
| Bagging | 0.819672 | 0.843750 | 0.818182 | 0.830769 | 0.819805 |
for t = 1,2,..,T:
Prediction:
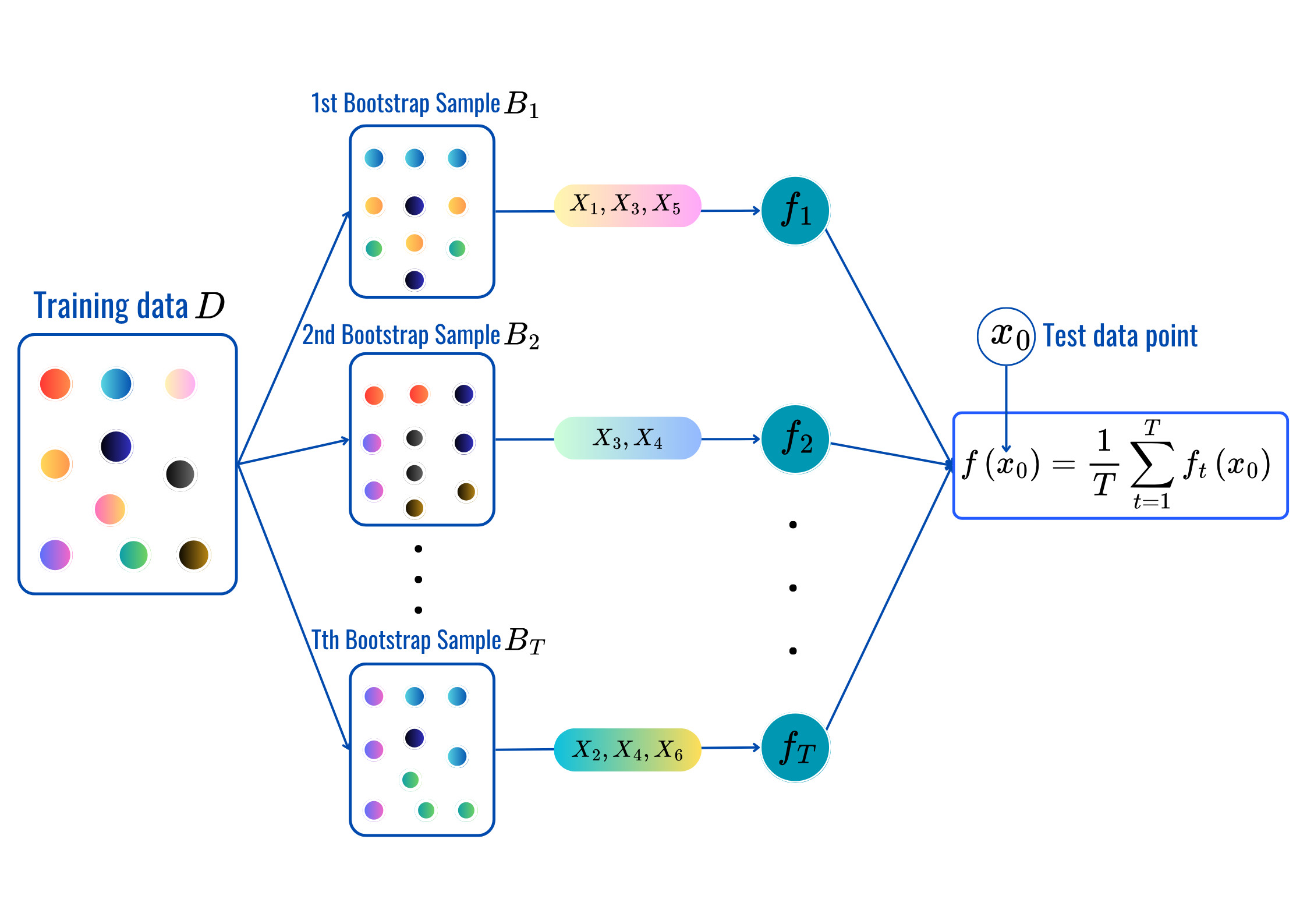
max_featuresmax_depth, min_sample_split, min_samples_leaf and citerion.from sklearn.ensemble import RandomForestClassifier
# Set up Bagging Classifier
rf = RandomForestClassifier()
# Define hyperparameter grid
param_grid = {
'n_estimators': [100, 200, 300, 500], # Number of trees in Bagging
'max_samples': [0.25, 0.4, 0.5, 0.6, 0.75, 0.9], # Proportion of samples per tree
'max_depth' : [5, 10, 20, 25, 30, 35, 40],
'criterion': ["gini", "entropy"], # Splitting criterion
}
grid_search = GridSearchCV(rf, param_grid, cv=5, n_jobs=-1, scoring='accuracy')
grid_search.fit(X_train_encoded, y_train)
# Best parameters and corresponding model
best_model = grid_search.best_estimator_
print(f"Best parameters: {grid_search.best_params_}")
# Make predictions with the best model
y_pred = best_model.predict(X_test_encoded)
test_perf = pd.concat([test_perf, pd.DataFrame(
data={'Accuracy': accuracy_score(y_test, y_pred),
'Precision': precision_score(y_test, y_pred),
'Recall': recall_score(y_test, y_pred),
'F1-score': f1_score(y_test, y_pred),
'AUC': roc_auc_score(y_test, y_pred)},
columns=["Accuracy", "Precision", "Recall", "F1-score", "AUC"],
index=["RF"])])
test_perfBest parameters: {'criterion': 'gini', 'max_depth': 25, 'max_samples': 0.75, 'n_estimators': 100}| Accuracy | Precision | Recall | F1-score | AUC | |
|---|---|---|---|---|---|
| LSVM | 0.852459 | 0.852941 | 0.878788 | 0.865672 | 0.850108 |
| RBF-SVM | 0.819672 | 0.866667 | 0.787879 | 0.825397 | 0.822511 |
| Poly3-SVM | 0.852459 | 0.875000 | 0.848485 | 0.861538 | 0.852814 |
| Bagging | 0.819672 | 0.843750 | 0.818182 | 0.830769 | 0.819805 |
| RF | 0.786885 | 0.812500 | 0.787879 | 0.800000 | 0.786797 |
for t = 1,2,..,T:
Prediction:
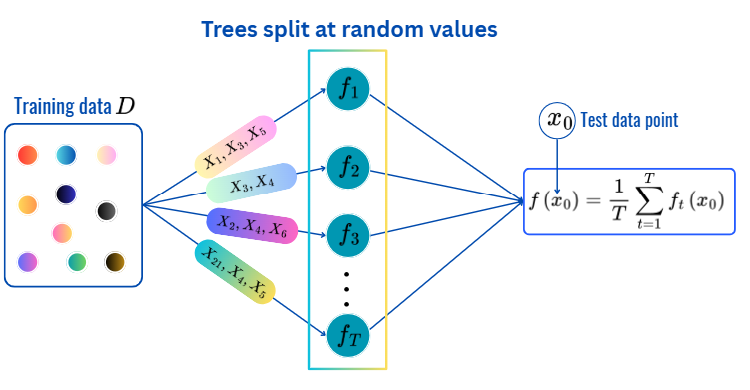
max_depth, min_sample_split, min_samples_leaf and citerion.from sklearn.ensemble import ExtraTreesClassifier
# Set up Bagging Classifier
extr = ExtraTreesClassifier()
# Define hyperparameter grid
param_grid = {
'n_estimators': [100, 200, 350, 500], # Number of trees in Bagging
'max_depth' : [5, 7, 10, 20, 30, 40],
'max_features' : [2, 3,5,7,9],
'criterion': ["gini", "entropy"], # Splitting criterion
}
grid_search = GridSearchCV(extr, param_grid, cv=5, n_jobs=-1, scoring='accuracy')
grid_search.fit(X_train_encoded, y_train)
# Best parameters and corresponding model
best_model = grid_search.best_estimator_
print(f"Best parameters: {grid_search.best_params_}")
# Make predictions with the best model
y_pred = best_model.predict(X_test_encoded)
test_perf = pd.concat([test_perf, pd.DataFrame(
data={'Accuracy': accuracy_score(y_test, y_pred),
'Precision': precision_score(y_test, y_pred),
'Recall': recall_score(y_test, y_pred),
'F1-score': f1_score(y_test, y_pred),
'AUC': roc_auc_score(y_test, y_pred)},
columns=["Accuracy", "Precision", "Recall", "F1-score", "AUC"],
index=["Extra-Trees"])])
test_perfBest parameters: {'criterion': 'gini', 'max_depth': 20, 'max_features': 3, 'n_estimators': 200}| Accuracy | Precision | Recall | F1-score | AUC | |
|---|---|---|---|---|---|
| LSVM | 0.852459 | 0.852941 | 0.878788 | 0.865672 | 0.850108 |
| RBF-SVM | 0.819672 | 0.866667 | 0.787879 | 0.825397 | 0.822511 |
| Poly3-SVM | 0.852459 | 0.875000 | 0.848485 | 0.861538 | 0.852814 |
| Bagging | 0.819672 | 0.843750 | 0.818182 | 0.830769 | 0.819805 |
| RF | 0.786885 | 0.812500 | 0.787879 | 0.800000 | 0.786797 |
| Extra-Trees | 0.836066 | 0.870968 | 0.818182 | 0.843750 | 0.837662 |
Each method relies on two main ideas:
The individual trees within each method may be too flexible (high variance), however averaging/voting results in more stable predictions.
There are many hyperparameters to be optimized (CV):
T: the number of treesfor t = 1, 2,..., T:
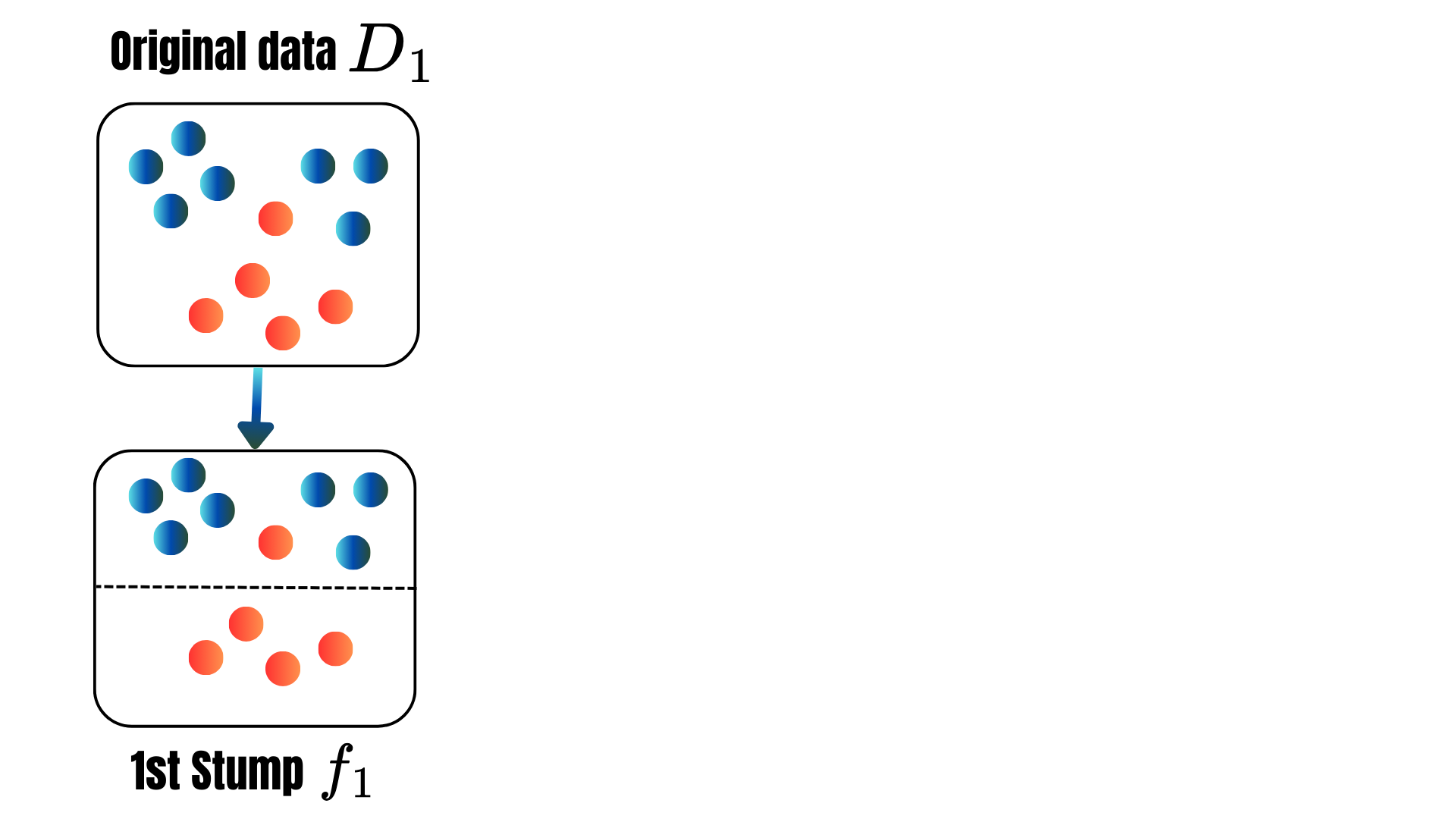
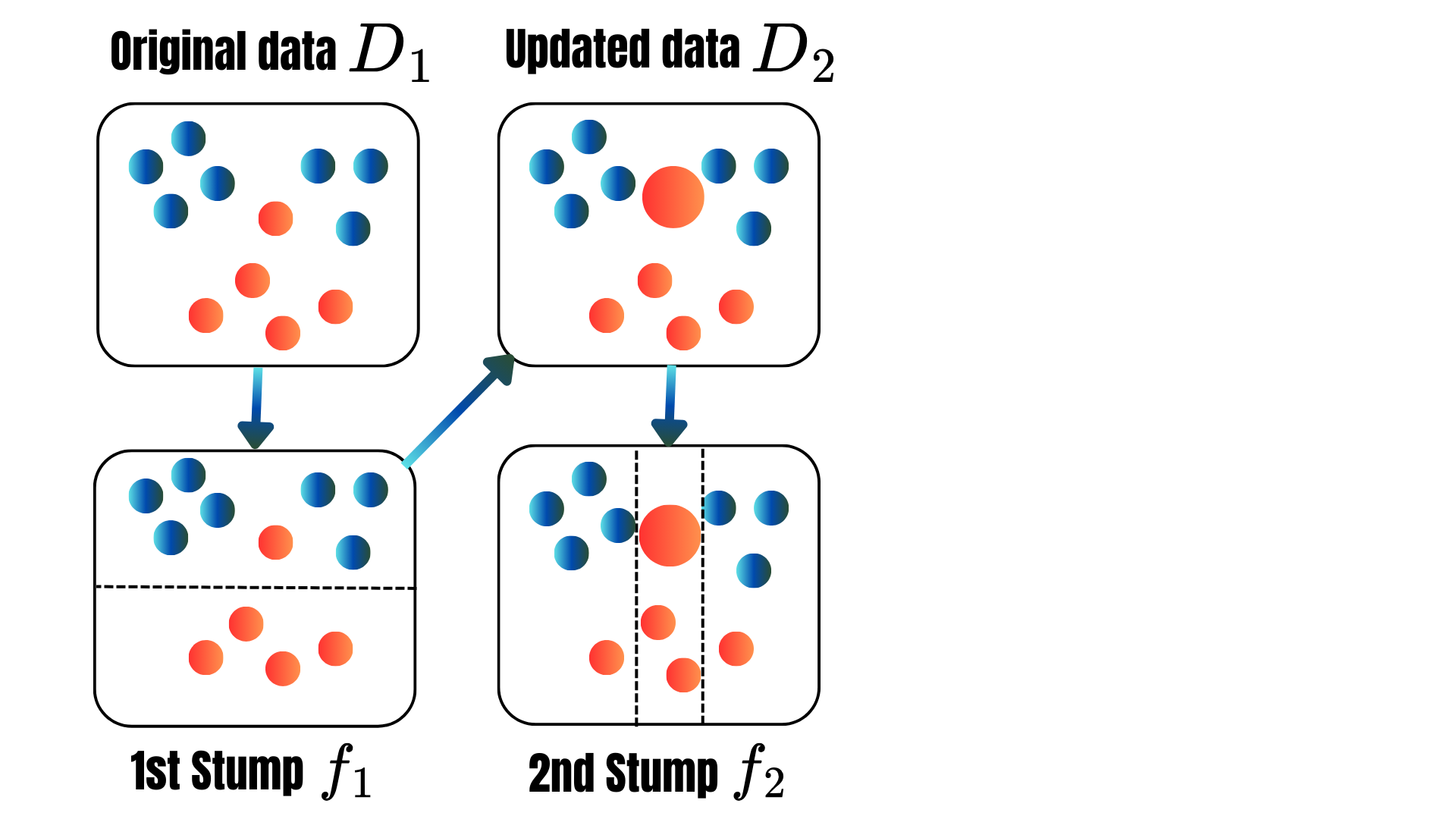
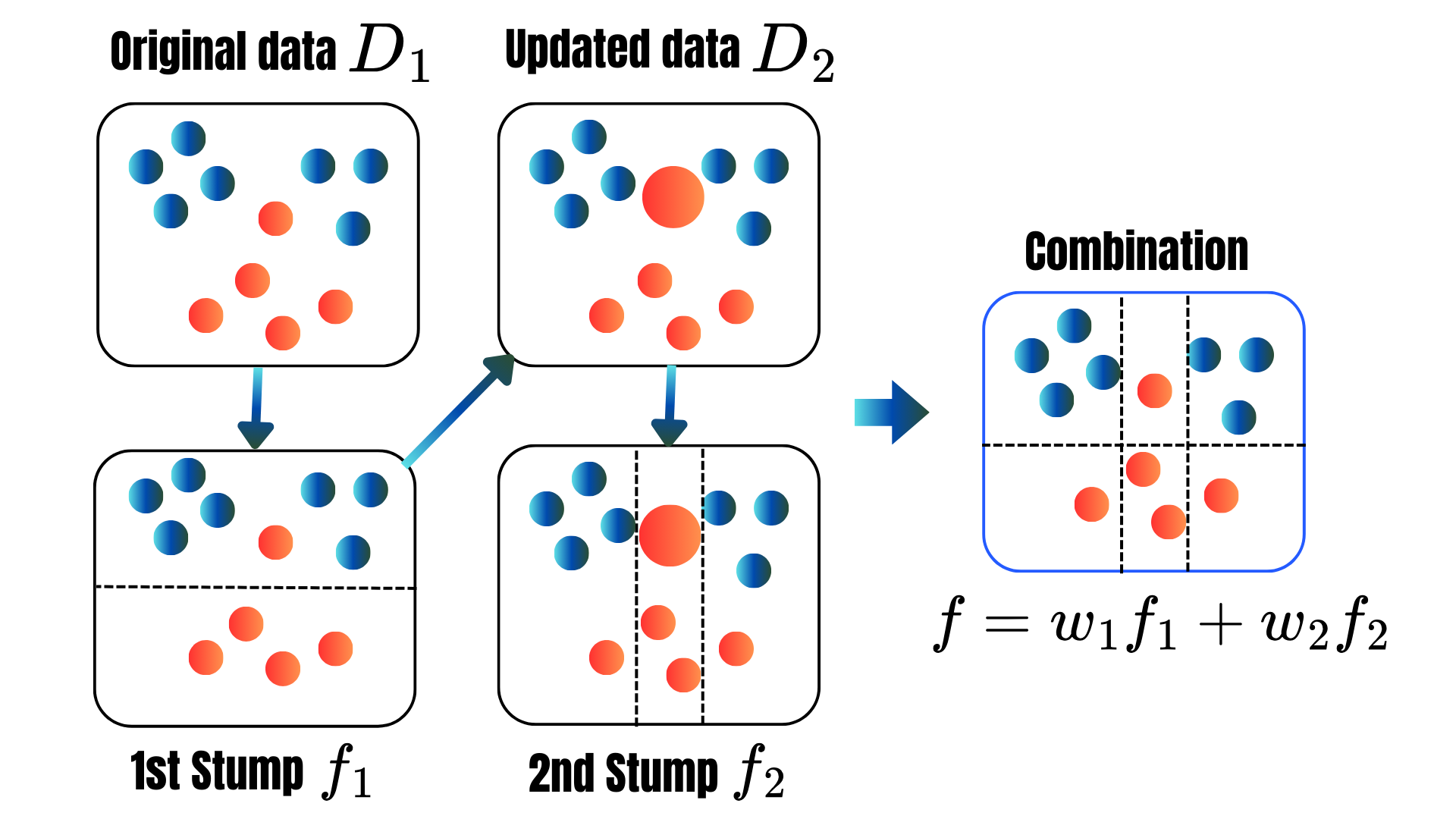
for t = 1, . . . , T:
for t = 1, 2, ..., M:
\[\begin{align*}L(F_t)&=\sum_{i=1}^nL(y_i,F_t(\text{X}_i))+\sum_{m=1}^t\Omega(f_m)\\ \Omega(f_t)&=\gamma T+\frac{1}{2}\lambda\|w\|^2.\\ L(F_t)&\approx \sum_{i=1}^n[g_iF_t(\text{x}_i)+\frac{1}{2}h_iF_t^2(\text{x}_i)]+\Omega(F_t),\\ g_i&=\frac{\partial L(y_i,F_t(\text{x}))}{\partial F_t(\text{x})}\|_{F_t(\text{x})=F_t(\text{x}_i)}\\ h_i&=\frac{\partial^2 L(y_i,F_t(\text{x}))}{\partial F_t(\text{x})^2}\|_{F_t(\text{x})=F_t(\text{x}_i)}. \end{align*}\]
to ensure the efficiency and accuracy of the method.
👉 Check out the notebook.
importances = best_model.feature_importances_
importance_df = pd.DataFrame({'Feature': list(X_train.select_dtypes(include="number").columns)+list(encoder.get_feature_names_out()), 'Importance': importances})
# Sort by importance
importance_df = importance_df.sort_values(by='Importance', ascending=False)
import plotly.express as px
# Plot feature importance using Plotly
fig = px.bar(importance_df, x='Feature', y='Importance', title='Feature Importance (Random Forest)',
labels={'Feature': 'Features', 'Importance': 'Importance Score'},
text=np.round(importance_df['Importance'], 3),
color='Importance', color_continuous_scale='Viridis')
fig.update_layout(xaxis_title="Features", yaxis_title="Importance Score", coloraxis_showscale=False, height=400, width=900)
fig.show()
